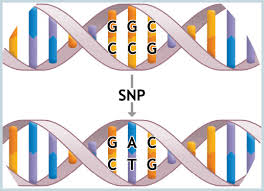
Single Nucleotide Polymorphism (SNP) Allele Frequency DNA Pools
6/3/2014
Lavebratt C, Sengul S. Single Nucleotide Polymorphism (SNP) Allele Frequency Estimation in DNA Pools using Pyrosequencing. Nat Protoc. 2006;1(6):2573-82.
Identifying the genetic variation underlying complex disease requires analysis of many single nucleotide polymorphisms (SNPs) in a large number of samples. Several high-throughput SNP genotyping techniques are available; however, their cost promotes the use of association screening with pooled DNA. This protocol describes the estimation of SNP allele frequencies in pools of DNA using the quantitative sequencing method Pyrosequencing (PSQ). PSQ is a relatively recently described high-throughput method for genotyping, allele frequency estimation and DNA methylation analysis based on the detection of real-time pyrophosphate release during synthesis of the complementary strand to a PCR product. The protocol involves the following steps: (i) quantity and quality assessment of individual DNA samples; (ii) DNA pooling, which may be undertaken at the pre- or post-PCR stage; (iii) PCR amplification of PSQ template containing the variable sequence region of interest; and (iv) PSQ to determine the frequency of alleles at a particular SNP site. Once the quantity and quality of individual DNA samples has been assessed, the protocol usually requires a few days for setting up pre-PCR pools, depending on sample number. After PCR amplification, preparation and analysis of PCR amplicon by PSQ takes 1 h per plate.